About me
I'm Abdullah Nayem Wasi Emran, currently working as a Lecturer at the Department of Computer Science and Engineering of
Brac University. I have completed my Bachelor's degree in Computer Science and Engineering from
Bangladesh University of Engineering and Technology (BUET).
My research sits at the intersection of AI/ML for Healthcare and Bioinformatics, with a focus on methods that remain reliable across shifts in data and population—particularly in low-resource settings.
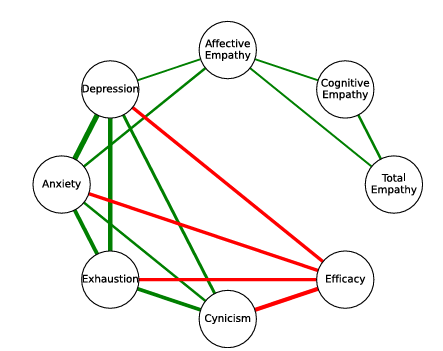
Recently, I led a cross-dataset study on mental health across diverse cohorts, published in
Humanities & Social Sciences Communications (Nature Portfolio, 2025). On the bioinformatics side, our
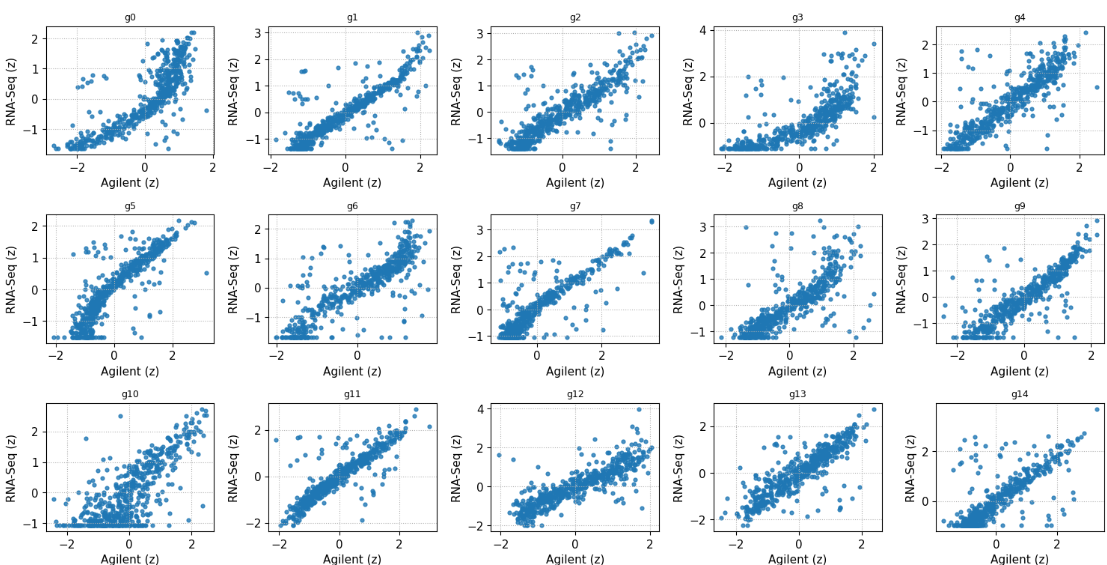
SCOPES work, accepted to the
AAAI-26 AI2ASE Workshop, proposes stability-aware, cross-platform feature selection for matched TCGA microarray/RNA-seq data—targeting reproducible biomarkers rather than fragile, platform-specific signals.
Broadly, I care about Causal and Fair Machine Learning: treatment-effect modeling, subgroup analysis, and evaluation protocols that reveal when a model’s apparent gains don’t translate to real-world impact.
My goal is to design simple, principled pipelines—data curation, robust learning, and transparent metrics—that can be deployed without exotic infrastructure.
Outside of academics, I enjoy learning about advances at the intersection of biology and computing.